Date: Wed, 25 Feb 2015 08:36:25 +0000
Hi George
Many thanks for your reply, I really appreciate it.
Thank you for confirming that there is no normalisation required to correct for the use of LAM-BIAS.
I have spent some time understanding the changes you suggested to my input file. I was interested in the idea of redefining the material with a lower density, increased length and subsequent smaller reduction factor in the LAM-BIAS. I hadn't thought of doing this before, neither had I seen it mentioned much in the discussion list.
I have spent time running your file (foil 10 cm, LAM-BIAS = 1e-2, 1e8 primaries) to compare to the results I had with my input file (foil one micron, LAM-BIAS = 5e-7, 50000 primaries). I was happy to see that for the most part they agreed remarkably well (see attached spreadsheet or picture). The main differences I observed were that initially my model created Boron-13 and at each decay time my model created Hydrogen-3. I tried modifying WHAT(5) of RADDECAY to see if this was the difference but it didn't seem to achieve agreement. Maybe it's a convergence issue? I wasn't able to run more than 1e8 particles in any useful time frame. Or maybe this is an artefact of the 'over-biasing' you mention in my input file? From my understanding the formation of tritium is important to consider.
Thank you once again for your email, it has certainly given me a better understanding of different ways of using FLUKA to solve problems, and given me some confidence that my input file was producing meaningful results.
Hayley
-----Original Message-----
From: owner-fluka-discuss_at_mi.infn.it [mailto:owner-fluka-discuss_at_mi.infn.it] On Behalf Of George Kharashvili
Sent: 19 February 2015 15:49
To: Smith, Hayley (STFC,RAL,ISIS)
Cc: FLUKA Discussion List
Subject: Re: [fluka-discuss]: RESNUCL with LAM-BIAS
Hello Hayley,
I think your foil may be too thin and your hadronic inelastic interactions - overbiased.
You can redefine your target material(s) (using MATERIAL card) with lower density and increase the target thickness so that you end up with the same g/cm^2. Using 1E-2 in WHAT(2) of LAM-BIAS is a reasonable value from my experience. I would also use WHAT(4) to define which particles the biasing applies to (even though the default may be protons).
RADDECAY - if you would like to kill both electromagnetic cascade and transport of decay radiation, set WHAT(5) of RADDECAY card to 9999999999.
There is no normalization needed to correct for the use of LAM-BIAS, FLUKA takes care of it all. However, the RESNUCLE units will be Bq/cc only if you assign the correct volume of the scoring region in WHAT(6). Otherwise it is Bq per region. I'm not sure why you are scoring RESNUCLE in all regions. Don't you wont to score it in the region called FOIL?
I have attached a modified version of your input file that you may find useful.
Cheers
- George
----- Original Message -----
From: "hayley smith" <hayley.smith_at_stfc.ac.uk>
To: "FLUKA Discussion List" <fluka-discuss_at_fluka.org>
Sent: Monday, February 16, 2015 11:44:29 AM
Subject: [fluka-discuss]: RESNUCL with LAM-BIAS
Hi
I am trying to look at the residual nuclei produced when a 70 MeV proton beam repeatedly passes through a thin (on the order of microns) foil. I am trying to understand the different residuals, and proportions of residuals, produced by different foil materials (carbon, aluminium etc) and I have a few questions about my methods:
From reading many of the previous FLUKA discuss posts I have understood that on these short length scales it is necessary to use LAM-BIAS to reduce the nuclear interaction length. At 70 MeV the .out file showed that the inelastic interaction length was 34.31 cm in carbon so I reduced the interaction length by 1e-6 using the LAM-BIAS card. Is this a reasonable scaling factor?
I am now looking at the RESNUCL output that I get but am confused about interpreting the results, even after consulting previous discussion list posts and the notes from the most recent course. I have tried repeating the simulation with LAM-BIAS for different numbers of primary particles (1000 - 50000) and different scaling factors to get an understanding. In my simulation I assume a proton distribution of 1.25e15 protons per second. I have looked at the .out files and .lis files but I am still unsure - do I need to apply any weighting to the RESNUCL output or has this already been taken into account when the Bq/cc are processed?
And are there any other settings that I may be missing in my very simple input file (attached) that would allow me to get a useful simulation of the residuals?
Many thanks for any clarification you are able to give,
Hayley
Hayley Smith
ISIS Facility
Rutherford Appleton Laboratory
Harwell Oxford
OX11 0QX
01235 445524
hayley.smith_at_stfc.ac.uk<mailto:hayley.smith_at_stfc.ac.uk>
__________________________________________________________________________
You can manage unsubscription from this mailing list at https://www.fluka.org/fluka.php?id=acc_info
- application/vnd.openxmlformats-officedocument.spreadsheetml.sheet attachment: Comparison.xlsx
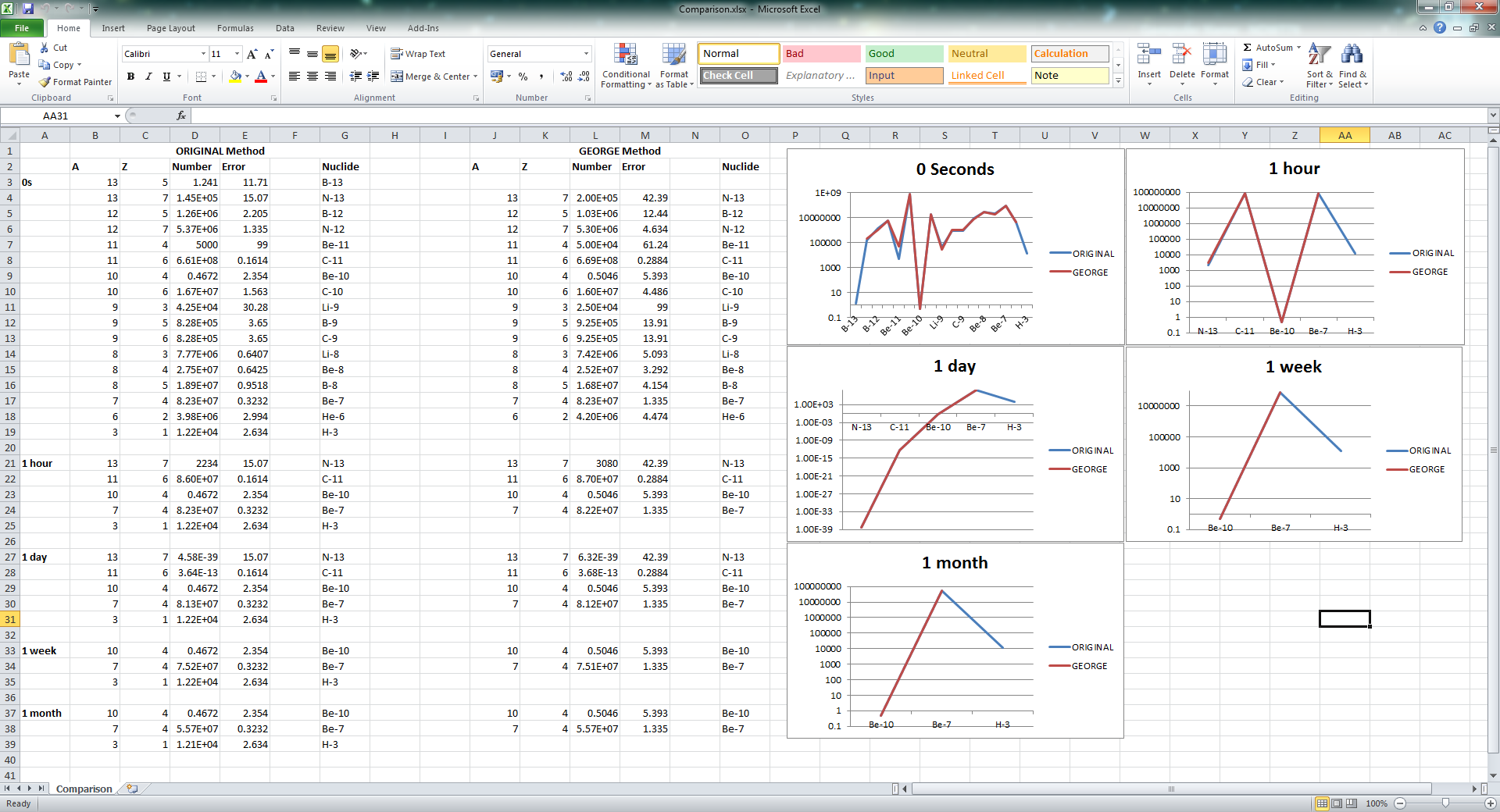
(image/png attachment: Comparison.png)